GWAS of Breast Cancer Subtypes Improves Understanding of Genetic Risk
, by DCEG Staff
A new genome-wide association study (GWAS) for breast cancer has identified susceptibility loci that increases our understanding of the genetic architecture of different breast cancer subtypes and expands the potential for the development of subtype-specific polygenic risk scores. The findings were published May 18, 2020, in Nature Genetics.
Montserrat García-Closas, M.D., Dr.P.H., and colleagues have been searching for genetic factors to explain breast cancer risk since the earliest genetic studies. To date, GWAS have revealed over 170 independent susceptibility loci, some of which have been linked to estrogen receptor (ER) status, an important predictor of disease outcome. However, other important tumor characteristics have yet to be well studied in relation to genetic predisposition. The latest report is the first to simultaneously account for multiple, correlated tumor markers to identify novel susceptibility loci and the specific sources of etiologic heterogeneity.
Investigators performed a GWAS including over 130,000 breast cancer cases and over 113,000 controls, plus over 18,000 BRCA1 mutation carriers (9,414 with breast cancer) of European ancestry. They identified 32 novel loci, 15 of which were associated with at least one tumor feature (ER, progesterone receptor (PR), human epidermal growth factor receptor 2 (HER2) status, or tumor grade). Five were linked to luminal- and non-luminal subtypes in opposite directions. Computational genomic analyses supported the observed differences in risk; the five loci contained cell-specific enhancers that differed by breast tissue cell type, normal luminal and basal mammary cells.
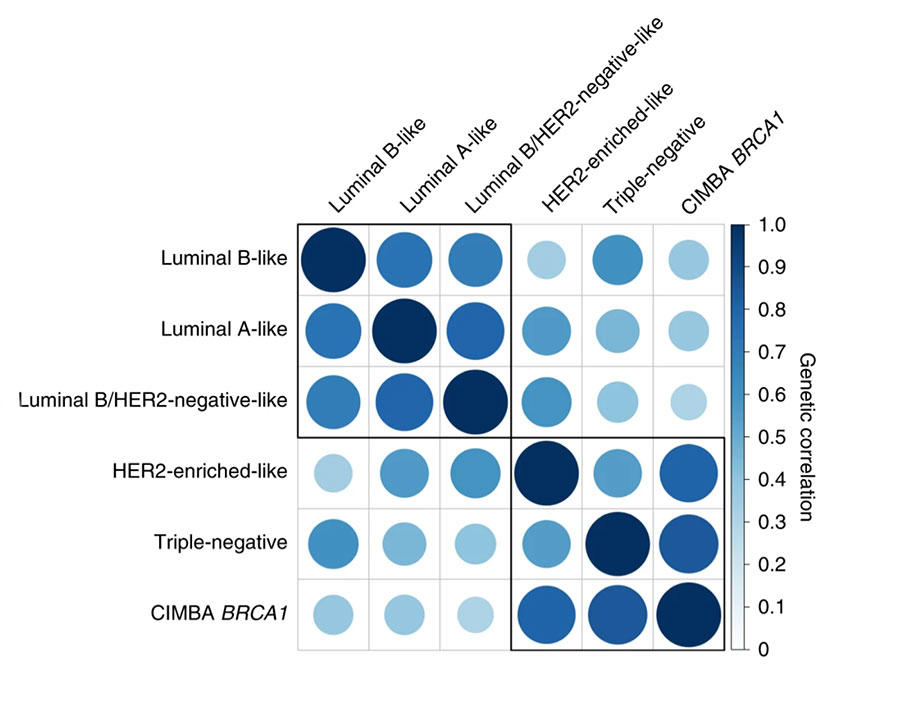
The investigators considered five subtypes defined by combinations of the tumor characteristics and found different levels of genetic correlation (i.e. sharing of genetic susceptibility loci), ranging from 0.35 to 0.80. Taken together, the 32 novel loci and those previously identified explained a higher proportion of the heritability due to common variants for luminal A-like (54.2%) than for triple-negative (37.6%) disease. Polygenic risk scores (PRSs) for different subtypes can identify 1% of women with over five-fold risk for luminal A-like, compared to women at average risk, and 1% of women at three-fold risk for triple-negative disease, compared to women at average risk.
In spite of its large sample size, the study faced important methodological challenges due to missing data on tumor characteristics and the high correlation between these characteristics. The investigators used novel statistical methods to address these challenges. Because the study population included only women of European ancestry, additional studies are needed to expand the generalizability of these findings for diverse populations.
Reference
Zhang, H. et al. Genome-wide association study identifies 32 novel breast cancer susceptibility loci from overall and subtype-specific analyses. Nat Genet (2020). DOI: 10.1038/s41588-020-0609-2