Genetic Architecture Study of 14 Cancers Describes Contribution of Common Variants
, by DCEG Staff
An international group of experts led by Montserrat García-Closas, M.D., Dr.Ph., and Dr. Nilanjan Chatterjee of the Johns Hopkins Bloomberg School of Public Health, analyzed data from multiple genome-wide association studies (GWAS) to estimate the number and effect size of common genetic variants associated with risk for 14 cancers and to describe their potential for risk stratification in populations of European ancestry. Their findings, which support the accumulating evidence that cancer is a highly polygenic disease, like other complex traits, were published July 2, 2020, in Nature Communications.
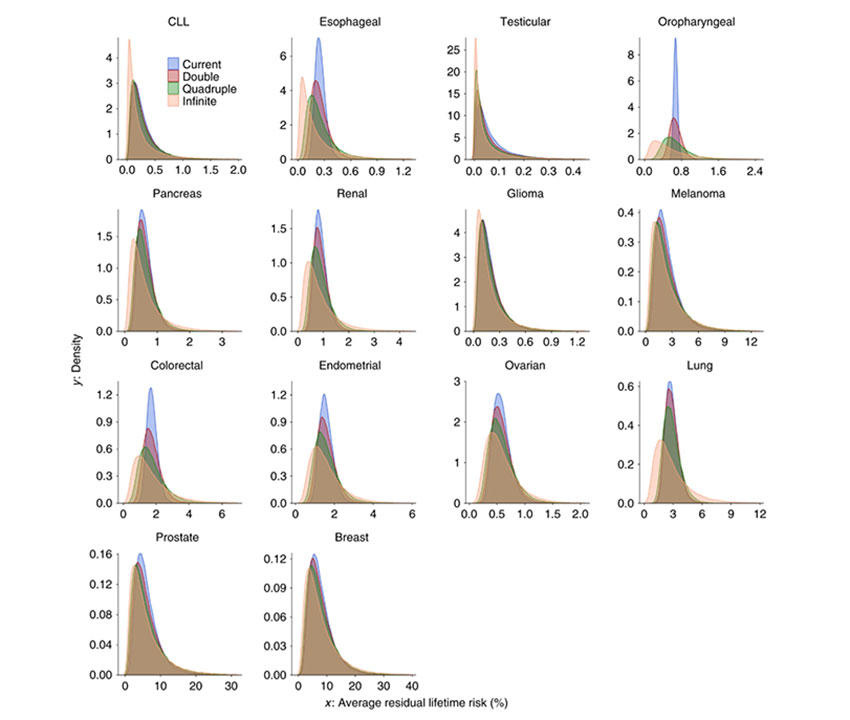
The cancers evaluated included breast, colorectal, endometrial, esophageal, glioma, lung, melanoma, oropharyngeal, ovarian, pancreas, prostate, renal, testicular, and chronic lymphocytic leukemia. All 14 showed a high degree of polygenicity that varied substantially across cancer types, involving—at a minimum— thousands of independent variants, from ~1,000 to 7,500.
The authors showed the strong potential of polygenic risk scores (PRS) for risk stratification for relatively common cancers, such as breast, colon, and prostate, but less so for malignancies with modest heritability and lower disease incidence. This reflects that large relative risks do not necessary translate into large degree of absolute risk stratification since this depends heavily on the underlying cancer incidence in the population. Clinical utility of PRS will further depend on the risk-reducing or early detection strategies available for a given cancer type.
To improve upon existing studies with currently available data, the authors project a wide range of sample sizes required to explain 80% of GWAS heritability—from 60,000 cases for testicular to over 1,000,000 cases for lung cancer.
By quadrupling the sample sizes of currently published GWAS, the percentage of GWAS heritability explained would rise to more than 40% across all cancers evaluated, except for oropharyngeal cancer. Such sample size increases would also lead to appreciable improvements in PRS discriminatory accuracy across all 14 malignancies.
Because the studies included only individuals with European ancestry, further studies are needed to evaluate if they are generalizable to a more diverse populations. The authors emphasize the need to amass studies with increasingly diverse populations.
Reference:
Zhang YD, et al. "Assessment of polygenic architecture and risk prediction based on common variants across fourteen cancers." Nature Commun 2020; doi.org/10.1038/s41467-020-16483-3.