Branch Profile: Biostatistics
, by Jennifer K. Loukissas, M.P.P.
Underpinning nearly every research study conducted by the Division of Cancer Epidemiology and Genetics (DCEG) is a collaboration with investigators in the Biostatistics Branch (BB).
Partners in Epidemiology
While highly trained and experienced epidemiologists can and do design their own analytic plans and calculate statistical power and other parameters, biostatisticians in BB ensure the statistical validity and strength of all DCEG research while actively engaging in the development of novel methods and tools. The mission of the branch is “to contribute to the understanding of cancer etiology and to improve public health through the development and application of quantitative methods.”
BB investigators approach their research in a number of different ways. “We form deep collaborative relationships with investigators across the Division and with scientists across the NCI, NIH, and throughout the scientific community,” said Paul S. Albert, Ph.D., Chief and senior investigator. “Analytic challenges in these scientific collaborations often provide the motivation for our independently initiated methodological research programs.”
Biostatisticians and statisticians lead projects that yield important discoveries in cancer etiology, risk prediction, and descriptive epidemiology, realized through state-of-the-art statistical and bioinformatic approaches. In addition to collaborating across the Division, BB investigators build partnerships within the branch, blending statistical expertise to solve challenging methodological problems in epidemiology and genetics.
To effectively explore questions in cancer etiology, biostatisticians in BB are proactive in the development of study designs that require the fewest resources possible, optimally utilize available biological samples and maximize power with the smallest study population needed.
Novel Methods for Emerging Fields of Study
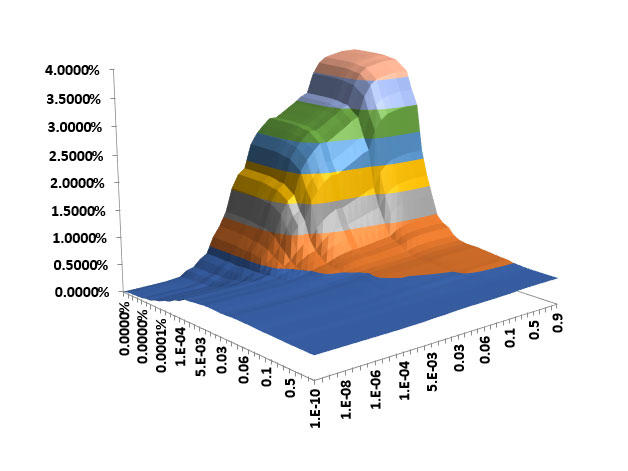
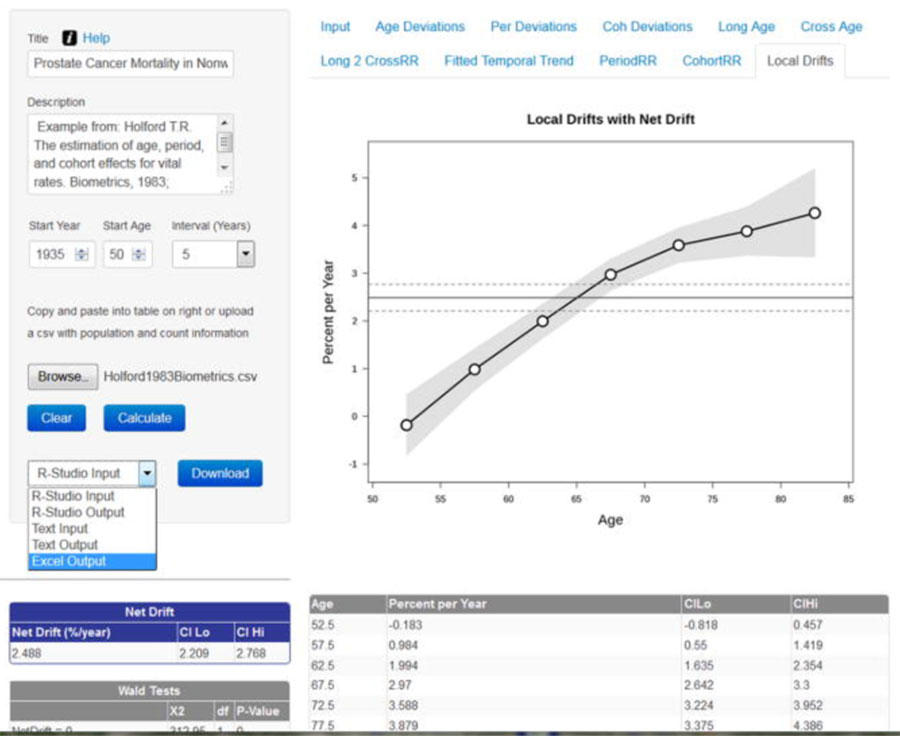
Over its history, BB has developed numerous tools for descriptive epidemiology and trends analysis that inform the direction of research across the Division. Phil S. Rosenberg, Ph.D., and colleagues perfected the Age Period Cohort analysis model and created a web-based application used widely for investigations of questions about breast cancer incidence by subtype, separate effect of the opioid epidemic on premature mortality, HIV-AIDS-associated cancers, and other outcomes.
Dozens of programs have been created and related methods published in the peer-reviewed literature to address emerging challenges in data analysis, particularly for molecular and genetic epidemiological studies. For example, Jianxian Shi, Ph.D., and Kai Yu, Ph.D., created analytic approaches to big data from genome-wide association (GWAS), whole genome, and exome sequencing studies. Together with the Integrative Tumor Epidemiology Branch, they came up with methods for analyzing tumor genomics data to reveal patterns of somatic mutations from exposures like cigarette smoking. Additionally, Dr. Yu has focused on pathway analysis for GWAS studies with investigators on a number of different cancer outcomes.
Biostatisticians respond to emerging research opportunities like metabolomics, microbiome, and tissue arrays, or novel risk factors by providing and testing nuanced approaches. For example, Joshua Sampson, Ph.D., and colleagues developed novel methods to identify metabolic profiles that are either predictive of disease or that can offer insight into why known risk factors are associated with disease.
Dr. Shi and Dr. Liu, in collaboration with MEB, created methods to analyze longitudinal microbiome data, quantifying the stability of the human microbiome over time. They are evaluating the impact of temporal variability in microbiome measurements on sample size requirements for etiologic studies. Results could have important implications for designing and analyzing future studies.
To test the performance of a new approach to capturing cancer incidence data for big cohorts, Dr. Liu compared questionnaire-based self-report of cancer diagnosis with registry incidence reports, confirming the value of the Virtual Pooled Registry Cancer Linkage System (VPR-CLS), which links research studies with U.S. cancer registries to ascertain incidence comprehensively through a secure online service.
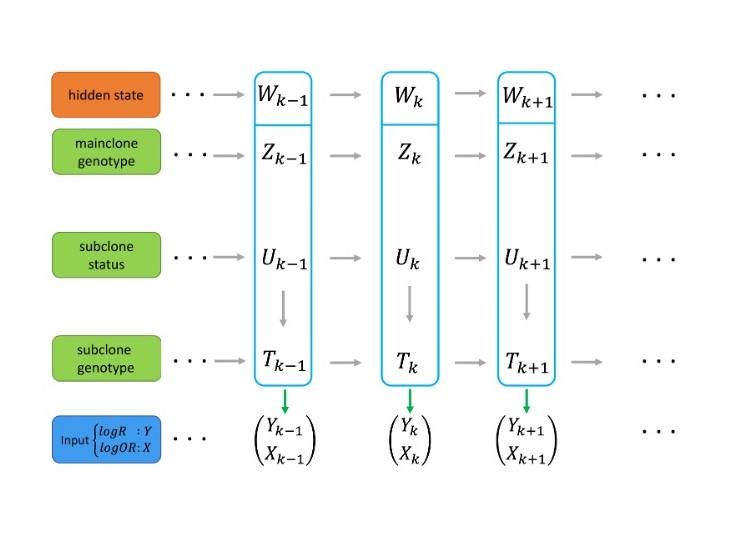
For example, Bin Zhu, Ph.D., and Dr. Albert used their respective expertise in somatic mutational analysis and hidden Markov modeling to develop novel methods for identifying copy number alterations in multiple clones among tumor samples (software package subHMM).
These tools and others are accessed via the branch web page.
Translating Research into Risk Prediction
In support of the DCEG mission, the branch focuses on new ways of studying existing or novel data to reveal patterns about cancer risk or etiology to inform public health prevention methods.
One of the first models to estimate a woman’s risk of developing breast cancer was published by Mitchell H. Gail, M.D., Ph.D., in the 1980s—known as the Gail Model, the NCI’s Breast Cancer Risk Assessment tool (BCRAT) provided important guidance for the enrollment of women into chemoprevention trials. In the decades since, Dr. Gail has updated BCRAT to incorporate data on minority women, and to reflect changes in population rates of breast cancer.
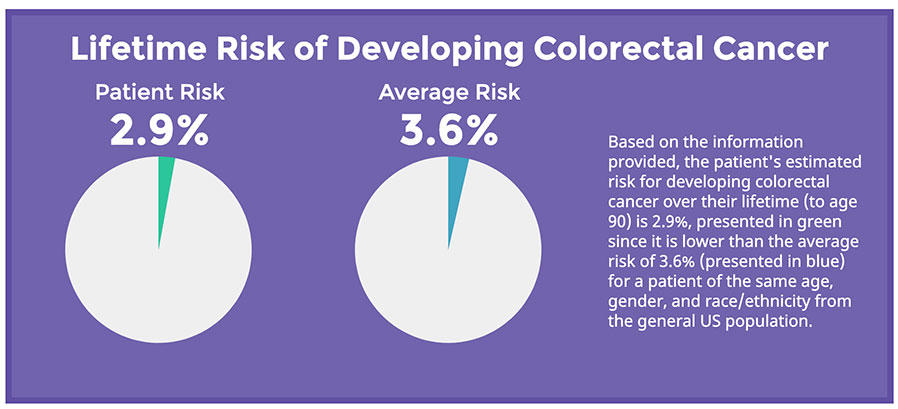
Ruth Pfeiffer, Ph.D., and colleagues published the first model to assess personalized risk for colorectal cancer. More recently, she developed and validated risk models for breast, endometrial, and ovarian cancer based on reproductive and lifestyle factors. Other efforts are underway facilitate the use of biomarkers like breast density, and polygenic risk scores in risk models. Tools like these are of tremendous interest to clinicians and the public who seek to better understanding how various factors influence disease risk.
Several investigators have designed statistical techniques to validate models for risk prediction or other investigations when important covariates are missing, such as exposure data, diagnostic information, such as tumor subtype, or other factors. These approaches allow epidemiologists to utilize imperfect data, and to inform the design of future studies.
A DCEG-wide team led by Hormuzd A. Katki, Ph.D., including Li C. Cheung, Ph.D., Rebecca Landy, and Corey Young, develops individualized prediction models to facilitate clinician-patient shared decision-making around lung cancer screening with low-dose computed tomography, based on the promising results of the NCI National Lung Cancer Screening Trial. In particular, they developed the first model to estimate individual increase in life-expectancy due to screening. Recent work focuses on how use of individual prediction models can reduce racial and ethnic disparities in screening eligibility.
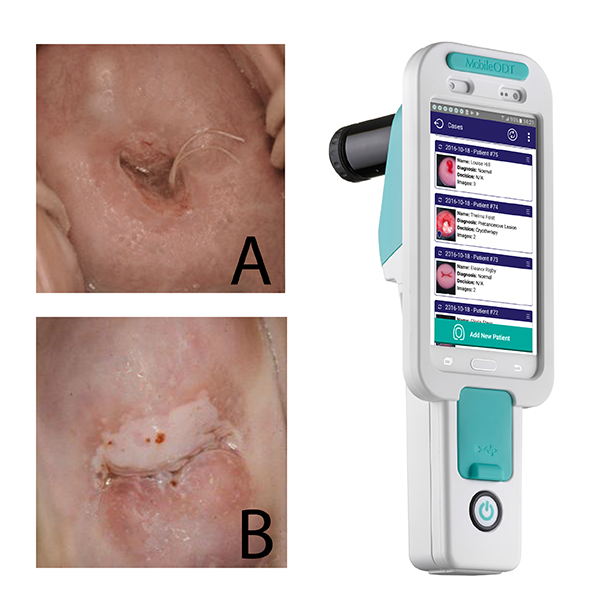
Numerous investigators in the branch have contributed to the accelerating the prevention of cervical cancer through studies of screening and efficacy and safety of the HPV vaccine with the Clinical Genetics Branch (CGB) and the Infections and Immunoepidemiology Branch (IIB). Dr. Yu collaborated with CGB and the National Library of Medicine on the development of an automated visual evaluation tool for cervical images to allow smart phones to be converted to a screening device for use in low-resource settings. Dr. Cheung collaborates with researchers in CGB to lead a team analyzing decades of screening data to use the risk of precancer/cancer to guide clinical actions that can interrupt the natural history of disease. Earlier this year, those methods were published alongside the 2019 ASCCP Risk-based Management Consensus Guidelines for abnormal cervical cancer screening results. In the arena of primary prevention, Dr. Sampson collaborates with IIB investigators on the design, monitoring, and analysis of HPV vaccine research, including ESCUDDO, the HPV one dose trial in Costa Rica.
Developing New Methodologies
A wide range of novel methods are in development, from approaches to analyzing cohort data collected from electronic medical records (EMR) to those for making inferences from epidemiologic cohort designs representative to the U.S. population. For example, to resolve differences in the make-up of prospective cohorts from a target population of interest, Barry I. Graubard, Ph.D.,and Dr. Katki proposed novel weighting methods to adjust cohort data to the U.S. population.
Also in development are new statistical methods for using longitudinal biomarkers for dynamic risk prediction. Drs. Liu, Albert, and colleagues are working to see whether longitudinal assessments of methylation markers can be used to dynamically assess the risk of cervical cancer or its precursors. Although these methods are being developed for HPV cohorts, they will play an important role in the Division’s Connect for Cancer Prevention Cohort Study, which will utilize repeated biomarkers, serial questionnaires, and EHR data.
In collaboration with the Occupational and Environmental Branch, Dr. Albert, Scientist Emeritus Jay H. Lubin, and colleagues are developing new methods for the study of environmental exposures on cancer risk. For example, most analyses within the Agricultural Health Study (AHS) have examined the effect of pesticide exposure one compound at a time. This segmented approach missed opportunities to more closely track the real-life exposure pattern of farmers. Although developed in the context of AHS, the methods will allow investigators throughout DCEG to study the simultaneous effects of a large number of exposures on cancer incidence. In related work, Dr. Albert collaborates with Mark Little, D.Phil., in the Radiation Epidemiology Branch to address measurement error from dosimetry in epidemiological studies of ionizing radiation exposure and cancer risk.
A Learning Culture: Fellowships, Training, and Mentoring
While maintaining its international reputation as a premier biostatistics research group, the branch is also home to a vibrant community of scholars. Dr. Albert prioritizes a collegial career-long learning culture, which builds technical expertise as well as relationships and morale. “We host weekly teas where staff discuss research but also build relationships. In the past year we have reinvigorated the seminar series for cross-training within the branch and across DCEG. More broadly, we maintain an active branch seminar series, featuring leading statisticians from across the U.S. Seminars are often attended by scientists throughout the Division, NCI, and NIH more generally.”
The Branch is currently home to more than 17 trainees, including pre- and postdoctoral fellows; five of whom hold joint training appointments with other DCEG branches. These cross-appointments reinforce the collaborative nature of the branch’s work, the critical role of BB across the Division’s research portfolio, and the focus on training that allows for development of expertise in exposures and outcomes as well as methods.
Numerous investigators have been recognized for their excellence in mentoring with awards from DCEG, the NCI and national professional associations.
Some trainees stay at the NCI in competitive tenure-track appointments within the branch, while others have been appointed to academic positions at major universities around the country, cancer centers, private industry, or continued in public health service at leading biostatistics groups at the NIH and FDA.
Research Service
In addition to the impressive research portfolio, the branch members are called upon to serve the scientific community in a number of capacities, on editorial boards of leading cancer and statistics journals, scientific planning committees for national and international professional societies, and Data Safety Monitoring Boards for important clinical trials throughout the NIH.
Many principal investigators in the branch publish their work not only in peer-reviewed journals, but in the academic press. For example, Drs. Pfeiffer and Gail published Absolute Risk: Methods and Applications in Clinical Management and Public Health, as part of the Monographs on Statistics and Applied Probability Series by CRC Press, drawing on their expertise and seminal achievements in modeling absolute risk.
Investigators serve on doctoral and mentoring committees for students as well as tenure-track investigators in DCEG. In addition, they are regularly invited to deliver seminars at major universities. In the last two years, seminars were given at Johns Hopkins, Harvard, University of Michigan, University of Pennsylvania, among others.
In reflecting on the value and expertise of the branch, Stephen J. Chanock, M.D., Director of DCEG, said, “what we have described above provides a snapshot of the impressive research accomplishments of the Biostatistics Branch. BB is integral to the success of the Division; each of us relies on our statistical colleagues for their keen insight, scientific rigor, energy, innovation, and dedication.”